An overview of the Hygrobates fluviatilis complex (Acariformes, Hydrachnidia, Hygrobatidae), with the description of a new species from Portugal and the first record of H. persicus from Cyprus
Pešić, Vladimir  1
; Girão, Dinis
1
; Girão, Dinis  2
; Tziortzis, Iakovos
2
; Tziortzis, Iakovos  3
; Papatheodoulou, Athina
3
; Papatheodoulou, Athina  4
and Ferreira, Sónia
4
and Ferreira, Sónia  5
5
1✉ University of Montenegro, Cetinjski put bb, 81000 Podgorica, Montenegro.
2CIBIO, Centro de Investigação em Biodiversidade e Recursos Genéticos, InBIO Laboratório Associado, Campus de Vairão, Universidade do Porto, 4485-661 Vairão, Vila do Conde, Portugal & Departamento de Biologia, Faculdade de Ciências, Universidade do Porto, 4169-007 Porto, Portugal & BIOPOLIS Program in Genomics, Biodiversity and Land Planning, CIBIO, Campus de Vairão, 4485-661 Vairão, Vila do Conde, Portugal.
3Water Development Department, Ministry of Agriculture, Rural Development and Environment, 100-110 Kennedy Avenue, 1047 Pallouriotissa, Cyprus.
4BIOMEHub Biomonitoring and Ecological Research, 4547, Mathikoloni- Limassol, Cyprus & Nature Conservation Unit, Frederick University, 1303, Nicosia, Cyprus.
5CIBIO, Centro de Investigação em Biodiversidade e Recursos Genéticos, InBIO Laboratório Associado, Campus de Vairão, Universidade do Porto, 4485-661 Vairão, Vila do Conde, Portugal & BIOPOLIS Program in Genomics, Biodiversity and Land Planning, CIBIO, Campus de Vairão, 4485-661 Vairão, Vila do Conde, Portugal.
2026 - Volume: 66 Issue: 2 pages: 327-340
https://doi.org/10.24349/j9sl-djd7ZooBank LSID: 77E60124-5B7D-4310-AC51-855DB73140B9
Original research
Keywords
Abstract
Introduction
Water mites of the Hygrobates fluviatilis complex represent one of the best documented groups for addressing biogeographical questions in cryptic water mite species (e.g., Pešić et al. 2017, 2019a, 2022). Over the last decade, several studies utilizing DNA barcoding data have revealed that Hygrobates fluviatilis, a name traditionally applied to specimens with a reticulated soft integument, actually consists of several distinct but previously overlooked lineages (see: Pešić et al. 2017, 2019b, 2020, 2023b).
Currently, the H. fluviatilis complex in the Western Palaearctic encompasses eleven species: Hygrobates fluviatilis (Ström, 1768); H. arenarius Smit & Pešić, 2017; H. corsicus Pešić & Smit, 2017; H. marezaensis Pešić & Dabert, 2017; H. turcicus Pešić, Esen & Dabert, 2017; H. persicus Pešić & Asadi, 2017; H. grabowskii Pešić, Saboori, Zawal & Dabert, 2019; H. ulii Pešić, Saboori, Zawal & Dabert, 2019; H. mediterraneus Pešić, 2020; H. balcanicus Pešić, 2020; and H. cyrnusensis Pešić & Smit, 2023. A time-calibrated phylogeny of this complex conducted by Pešić et al. (2023a) revealed that the most recent common ancestor originated during the Oligocene. Divergence between the main lineages appears to have occurred in the early Miocene (ca. 24 Ma), with the H. turcicus clade representing the earliest branching lineage.
In this paper, we provide an overview of the species of the Hygrobates fluviatilis complex using publicly available data from the Barcode of Life Data (BOLD) Systems and from GenBank. By analyzing species distribution patterns based on their Barcode Index Number (BIN) affiliations, we aim to offer new insights that stimulate future taxonomic and biogeographic research. Furthermore, as a continuation of our recent studies on Portuguese water mites (e.g., Pešić et al. 2024a, b, 2025d, 2026a) conducted within the framework of the Biodiversity Genomics Europe (BGE) project, a new species of the H. fluviatilis complex is described herein.
Material and methods
Water mites were collected using hand nets, sorted live in the field, and immediately preserved in 96% ethanol. Specimens intended for molecular analysis were examined without dissection under a stereo microscope in ethanol. Following DNA extraction, selected specimens were dissected and slide-mounted in Faure's medium.
Morphological nomenclature follows Gerecke et al. (2016); for detailed explanations concerning the morphology and measurements of the H. fluviatilis complex, see Pešić et al. (2017: Figure 1B–D). The genital acetabula in both sexes and the female genital plates were measured on both sides; thus, the dimensions are presented as ranges. The holotype and paratypes of the new species are deposited in the Naturalis Biodiversity Center, Leiden (RMNH).
All measurements are in µm. The following abbreviations are used: Ac-1 to -3 = acetabula from anterior to posterior; Cx-I to Cx-IV = coxae I to IV; dL = dorsal length; H = height; I-L-4-6 = fourth to sixth segments of first leg; L = length; Ma (mega-annum) = million years; mL = median length; n = number of specimens examined; P-1 to P-5 = palp segments 1 to 5; W = width.
Molecular analysis
Molecular analysis was conducted partly at the Canadian Centre for DNA Barcoding (Guelph, Ontario, Canada, CCDB; http://ccdb.ca/ ![]() ), partly at the University of Florence (UNIFI). DNA was extracted using a non-destructive protocol. For the methods used for cytochrome c oxidase subunit I (COI) gene amplification and sequencing, see Pešić et al. (2025b). Raw reads were demultiplexed using the Pacific Biosciences SMRT Link software. Consensus sequences were generated with the PacBio Amplicon Analysis (pbaa) tool. Primer trimming, translation and stop codon checking were performed using Geneious Prime 2024.0.1. Consensus sequences are made available in the Barcode of Life Data Systems (BOLD). Relevant voucher information, photos, and recently generated DNA barcodes are publicly accessible through the Dataset - DS-BGEPL06 (BGE Biodiversity Genomics Europe - Molecular phylogeny of Hygrobates fluviatilis complex) in BOLD. In total, DNA was extracted from six specimens of the genus Hygrobates from Portugal and six specimens of H. persicus from Cyprus (Table 1). For all other species, COI sequence data were obtained from Pešić et al. (2017, 2019b, 2020, 2021a, b, 2023a, b, c, 2024a, 2025a, c, 2026b), Pešić & Smit (2022), and Klimov et al. (2022) via the respective sequence data archives. In total, 325 sequences representing 12 species of the Hygrobates fluviatilis complex were analyzed, with Hygrobates prosiliens from the Netherlands used as the outgroup.
), partly at the University of Florence (UNIFI). DNA was extracted using a non-destructive protocol. For the methods used for cytochrome c oxidase subunit I (COI) gene amplification and sequencing, see Pešić et al. (2025b). Raw reads were demultiplexed using the Pacific Biosciences SMRT Link software. Consensus sequences were generated with the PacBio Amplicon Analysis (pbaa) tool. Primer trimming, translation and stop codon checking were performed using Geneious Prime 2024.0.1. Consensus sequences are made available in the Barcode of Life Data Systems (BOLD). Relevant voucher information, photos, and recently generated DNA barcodes are publicly accessible through the Dataset - DS-BGEPL06 (BGE Biodiversity Genomics Europe - Molecular phylogeny of Hygrobates fluviatilis complex) in BOLD. In total, DNA was extracted from six specimens of the genus Hygrobates from Portugal and six specimens of H. persicus from Cyprus (Table 1). For all other species, COI sequence data were obtained from Pešić et al. (2017, 2019b, 2020, 2021a, b, 2023a, b, c, 2024a, 2025a, c, 2026b), Pešić & Smit (2022), and Klimov et al. (2022) via the respective sequence data archives. In total, 325 sequences representing 12 species of the Hygrobates fluviatilis complex were analyzed, with Hygrobates prosiliens from the Netherlands used as the outgroup.
Download as
Locality
Coordinates
Process ID
Voucher ID
BIN BOLD
Hygrobates lusitanicus sp. nov.
Portugal, Braga, Tregosa, Azenha das Poldras
41.622° N, 8.712° W
DTBIO2366-25
BGPT_018_H02
BOLD:AFX1756
Portugal, Porto, Lousada, Moinho da Tapada
41.263° N, 8.307° W
BBIOP063-24
BGE_00109_F03
Portugal, Bragança, Mirandela, Torre de Dona Chama
41.665° N, 7.146° W
BBIOP028-24
BGE_00109_C04
BBIOP014-24
BGE_00109_B02
BBIOP029-24
BGE_00109_C05
Portugal, Bragança, Vinhais, Gasparona
41.850° N, 7.013° W
BBIOP037-24
BGE_00109_D01
Hygrobates persicus
Cyprus, Limassol, Mesa Potamos
34.876° N, 32.910° E
CYPTU010-25
CCDB_45908_A10
BOLD:ACB5533
CYPTU018-25
CCDB_45908_B06
CYPTU027-25
CCDB_45908_C03
CYPTU031-25
CCDB_45908_C07
Cyprus, Limnatis (Zygos) stream nr Kouris Dam
34.786° N, 32.941° E
CYPTU034-25
CCDB_45908_C10
Cyprus, Kochinas river
34.877° N, 32.910° E
CYPTU089-25
CCDB_45908_H05
Sequence alignments were performed using MUSCLE (Edgar 2004). Intra- and interspecific genetic distances were calculated based on the Kimura 2-parameter (K2P) model (Kimura 1980) using MEGA-X (Kumar et al. 2018). Additionally, sequence data were analyzed using Assemble Species by Automatic Partitioning (ASAP), a hierarchical clustering algorithm for species partitioning based on pairwise distance distributions (Puillandre et al. 2021). We utilized the online ASAP interface (https://spartexplorer.mnhn.fr/ ![]() ) with default settings and the K2P distance model.
) with default settings and the K2P distance model.
Results
Species delimitation in the Hygrobates fluviatilis complex using DNA-barcodes
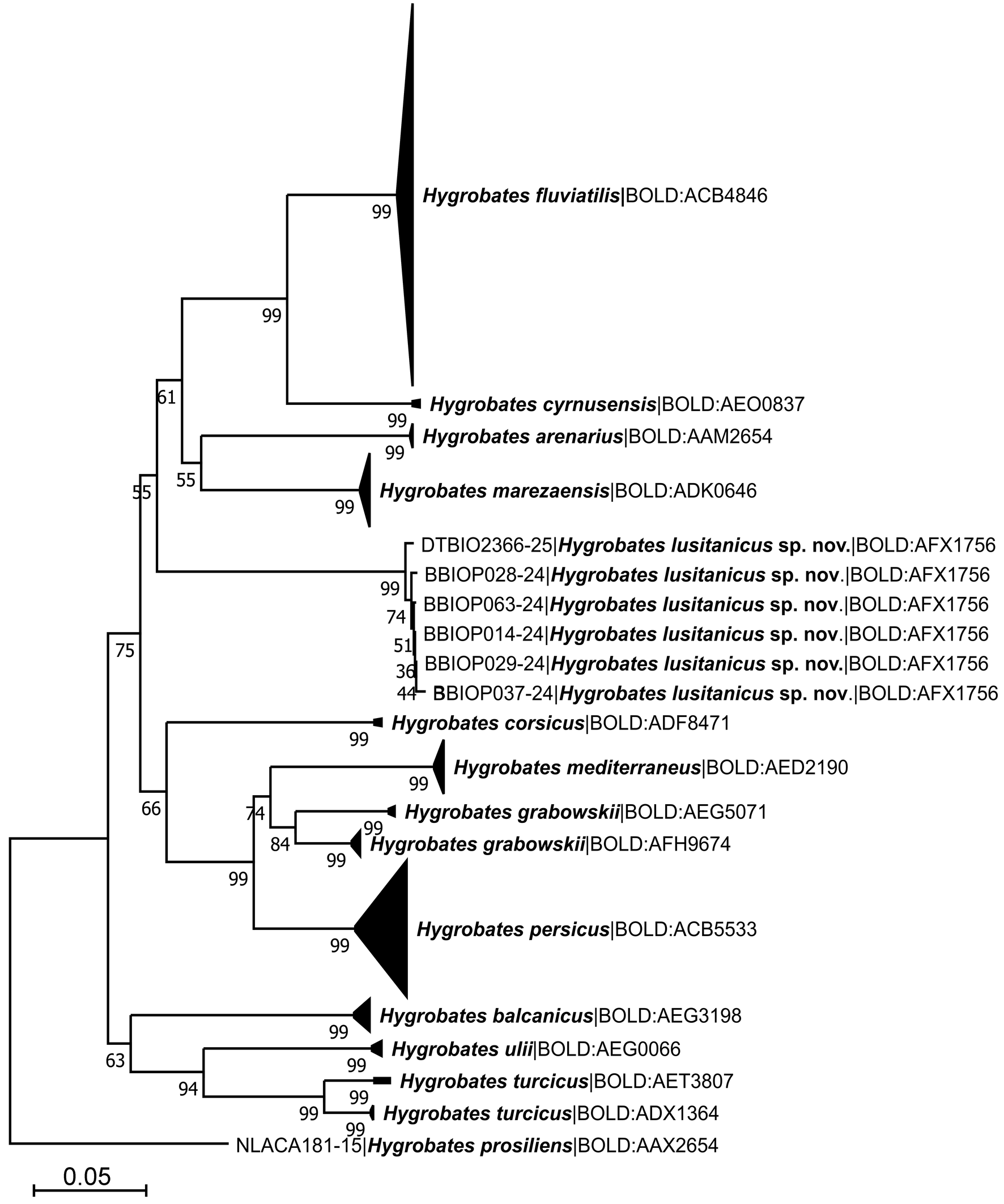


The final alignment for species delimitation using COI sequence data comprised 1,222 nucleotide positions (bp) for 325 specimens of the H. fluviatilis complex, with one specimen of H. prosiliens used as an outgroup to root the tree. The nucleotide sequences were translated into amino acid sequences without any observed stop codons. The Neighbor-Joining (NJ) tree is presented in Figure 1.
Download as
Species
Intragroup
Intergroup
-1
-2
-3
-4
-5
-6
-7
-8
-9
-10
-11
(1)* Hygrobates arenarius*
0.0006
(2) Hygrobates balcanicus
0.0103
0.258
(3) Hygrobates corsicus
0.0032
0.225
0.261
(4) Hygrobates cyrnusensis
0.0041
0.213
0.238
0.24
(5) Hygrobates fluviatilis
0.0153
0.208
0.256
0.229
0.117
(6) Hygrobates grabowskii
0.0345
0.23
0.236
0.202
0.204
0.224
(7) Hygrobates lusitanicus sp. nov.
0.0046
0.22
0.267
0.235
0.25
0.231
0.236
(8) Hygrobates marezaensis
0.0062
0.175
0.249
0.233
0.179
0.192
0.219
0.222
(9) Hygrobates mediterraneus
0.0082
0.249
0.264
0.211
0.247
0.267
0.129
0.228
0.221
(10) Hygrobates persicus
0.013
0.218
0.27
0.198
0.216
0.234
0.118
0.233
0.211
0.145
(11) Hygrobates turcicus
0.0216
0.279
0.197
0.252
0.235
0.256
0.226
0.276
0.24
0.241
0.247
(12) Hygrobates ulii
0.0057
0.268
0.239
0.249
0.261
0.263
0.246
0.266
0.248
0.24
0.259
0.154
Phylogenetic analyses clustered the COI sequences of the analyzed Hygrobates specimens into 13 strongly supported clades (Figure 1), eleven of which corresponded to previously known members of the H. fluviatilis complex. The COI sequences from the Hygrobates specimens collected in Portugal recovered as a sister clade to the H. fluviatilis group, albeit with moderate bootstrap support (55%). The H. fluviatilis group, a clade defined within the H. fluviatilis complex by Pešić et al. (2023), comprises H. fluviatilis, H. marezaensis, H. arenarius, and H. cyrnusensis. The genetic distance between H. lusitanicus sp. nov. and other species within the H. fluviatilis group ranged from 22.0% (with H. arenarius) to 25.0% K2P (with H. cyrnusensis) (see Table 2). These distances significantly exceed the barcoding gap identified by the ASAP method (6–10%) among all species within the H. fluviatilis complex (Figure 2), providing further evidence for the specific status of the new species from Portugal.
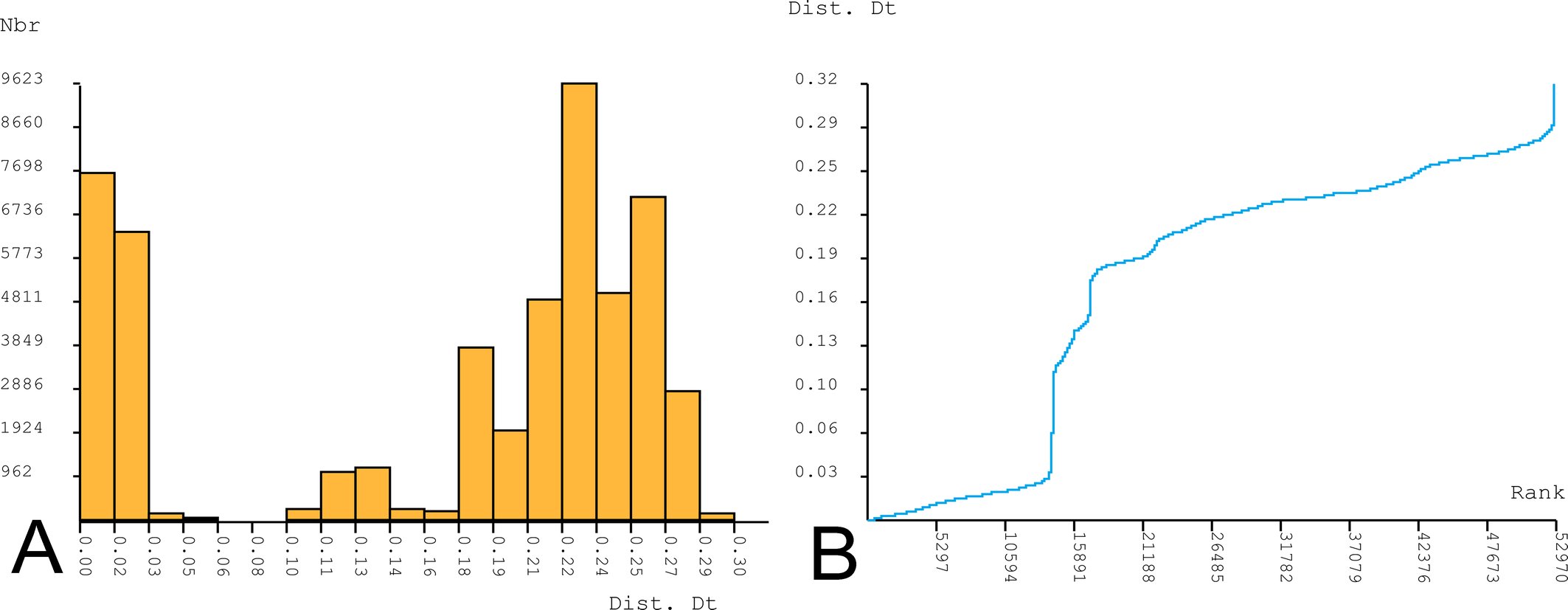


Systematic descriptions
Family Hygrobatidae Koch, 1842
Genus Hygrobates Koch, 1837
Subgenus Hygrobates s.s.
Hygrobates lusitanicus Pešić & Girão sp. nov.
ZOOBANK: 73E36162-125F-40B1-A884-CE117EA2C048 ![]()
Figures 3-4
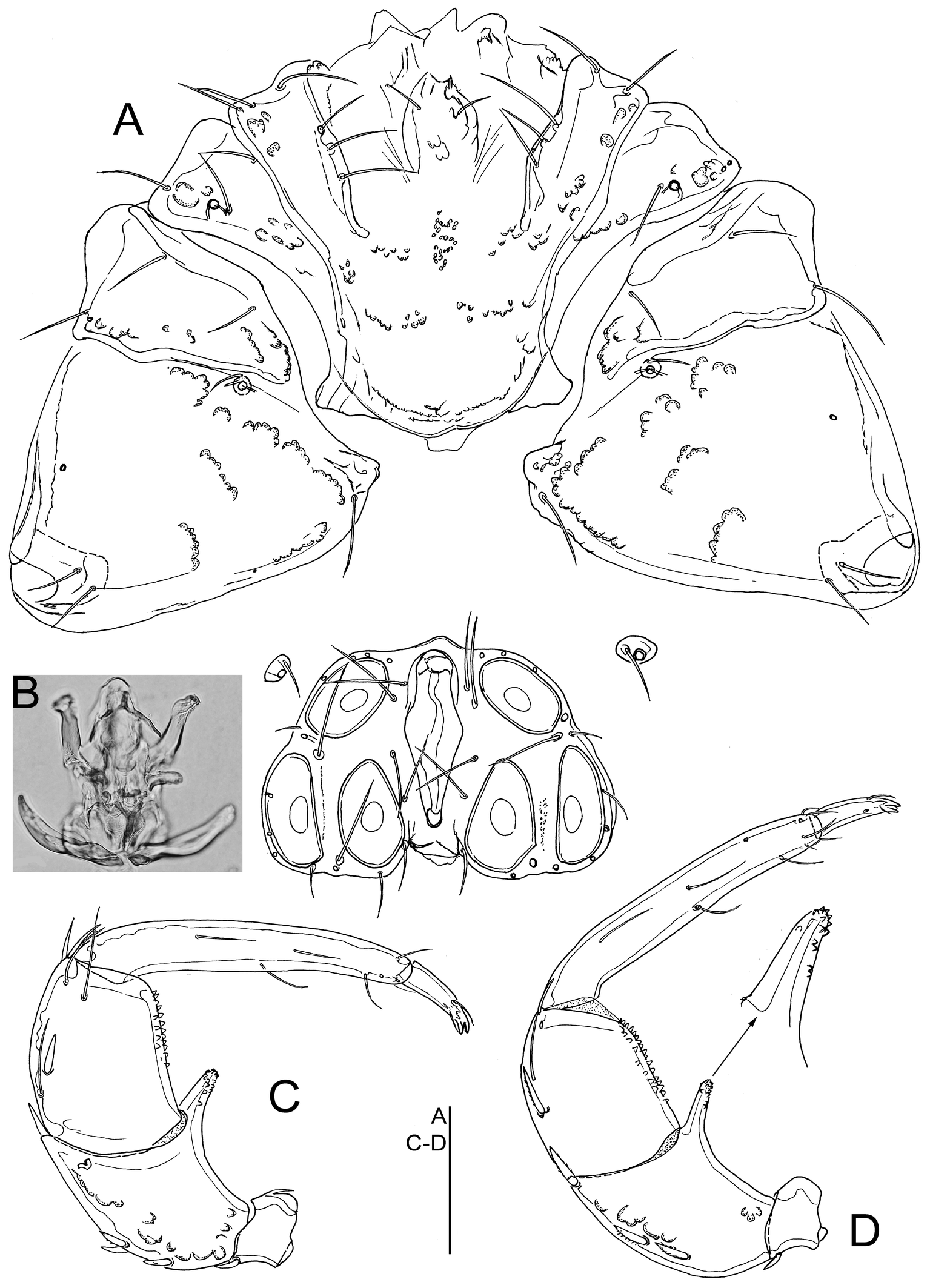


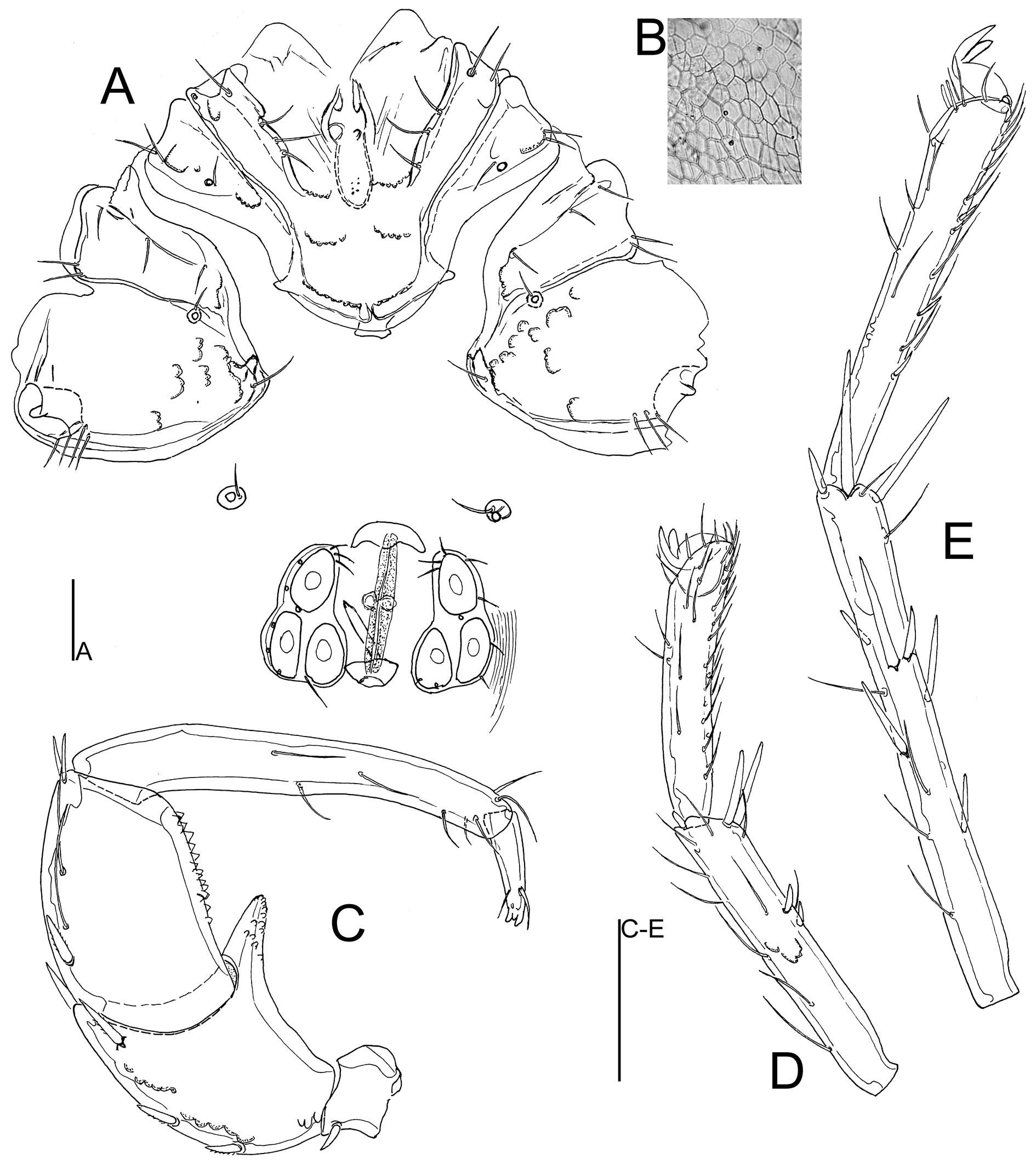


Material examined — Holotype ♂, sequenced (Voucher Id: BGE_00109_C04; BOLD Id: BBIOP028-24), dissected and slide mounted (RMNH), Portugal, Bragança, Mirandela, Torre de Dona Chama, Tuela river (Figure 5C), 41.665°N, 7.146°W, 254 m a.s.l., 13 July 2023, leg. Ferreira & Padilha. Paratypes: 2♀, same data as holotype, 1♀ dissected and slide mounted (RMNH).
Additional material – Portugal, Braga, Tregosa, Azenha das Poldras, Neiva river, 41.622°N, 8.712°W, 28 m a.s.l., 13 April 2023, leg. Ferreira, Girão & da Silva, 1♂.
Diagnosis — Membranous integument dorsally reticulate; postero-medial margin of Cx-I+II broadly rounded with well-developed subcutaneous extensions at the posterior margin of Cx-I; P-2 ventrodistal projection long and slender, denticulation restricted to the projection; P-4 L/H ratio 5.8–5.9; the female gonopore shorter than the genital plates (L ratio gonopore/genital plate < 1.0).
Description. General features — Colour yellowish to dark brown. Membranous integument dorsally reticulate. Postero-medial margin of Cx-I+II convex with well-developed subcutaneous extensions at the posterior margin of Cx-I (Figure 3A); Cx-IV subtriangular, with a distinctly nose-like protruding medial margin. Genital field with acetabula in a triangular arrangement. P-2 ventral margin almost straight and without denticulation, ventrodistally protruding in a long and slender, finger-like projection which is ventrally apically covered by denticulation (Figure 3C–D) P-3 with denticles covering distal two thirds of ventral margin. Male — Anterior margin of genital field convex with a knob-shaped medial projection, posterior margin deeply indented, with a central protrusion not extending beyond posterior genital plate margin (Fig. 3A). Female — Gonopore shorter than the genital plates (Figure 4A); P-4 more slender than in male.
Measurements. Male (holotype; BGE 00109 C04) — Idiosoma L 795, W 612; coxal field: L 388; Cx-III W 500; mL of Cx-I + gnathosoma 294; distance between lateralmost ends of Cx-II apodemes, 172. Genital field L/W 167/234, ratio; L Ac-1-3: 66, 78-81, 75-78. Ejaculatory complex L 150.
Palp — Total L 589; dL/H, dL/H ratio: P-1, 31/48, 0.65; P-2, 150/86, 1.7; P-3, 124/91, 1.4; P-4, 225/38, 5.9; P-5, 59/19, 3.1; P-2/P-4 ratio 0.67. Chelicera total L 306.
Legs — dL of I-L-1–6: 73, 100, 137, 194, 206, 190; dL of IV-L-1–6: 161, 163, 219, 322, 347, 272.
Female (paratype) — Idiosoma L 1180, W 950; coxal field: L 547; Cx-III W 682; mL of Cx-I + gnathosoma 383; distance between lateralmost ends of Cx-II apodemes, 219. Genital field L/W 209/288; genital plate L 175-180; gonopore L 169; L gonopore/genital plate ratio 0.94-0.97; L-Ac 1-3: 75-78, 81-89, 72-73.
Palp — Total L 706; dL/H, dL/H ratio: P-1, 34/54, 0.6; P-2, 172/103, 1.7; P-3, 155/99, 1.6; P-4, 273/49, 5.8; P-5, 72/21, 3.4; P-2/P-4 ratio 0.63. Chelicera total L 364, L basal segment 251, L claw 126, L basal segment/claw ratio 2.0.
Legs — dL of I-L-1–6: 84, 118, 148, 214, 228, 213; dL of IV-L-1–6: 163, 181, 237, 360, 372, 294.
Etymology — The name refers to the geographical provenance (from Latin lusitanicus, Lusitanian, Portuguese) from which the new species is currently known.
Discussion — Phylogenetic analysis based on COI data placed Hygrobates lusitanicus sp. nov. as a sister clade to the fluviatilis-group, which comprises H. fluviatilis, H. marezaensis, H. arenarius, and H. cyrnusensis (see Pešić et al. 2023a). The level of COI differentiation between H. lusitanicus sp. nov. and these species is high (\textgreater22% K2P divergence), suggesting a long, independent evolutionary history for the new species.
Morphologically, among the above listed species the new species most closely resembles H. arenarius. Both species share a comparatively long, slender, finger-like distoventral projection on P-2, with ventral denticulation restricted to this projection. However, H. arenarius can be distinguished from H. lusitanicus sp. nov. by the following features: (1) the posteromedial margin of Cx-I+II in both sexes is more triangular, lacking the well-developed subcutaneous extension on the posterior margin of Cx-I present in the new species; (2) a slenderer P-4, with an L/H ratio of 6.7–7.8 (compared to 5.8–5.9 in H. lusitanicus sp. nov.); and (3) the female gonopore being distinctly longer than the genital plates (L ratio gonopore/genital plate > 1.0).
Distribution — Portugal: Braga, Bragança and Porto.
An overview of the Hygrobates fluviatilis complex in the Western Palaearctic
1. Hygrobates fluviatilis (Strøm, 1768)
Acarus fluviatilis Ström, 1768: 349.
Type locality — Norway: Heltneelva (Ström 1768); neotype designated by Pešić et al. (2017) from Møre og Romsdal Volda, Heltneelva ved Årneset (62.145°N, 6.097°E).
BIN — BOLD:ACB4846
BOLD Records — 167 [238 Private].
BIN Distribution — Bosnia and Herzegovina, Croatia, Germany, Montenegro, The Netherlands, Norway, Portugal, Romania, Russia, Serbia, Sweden.
2. Hygrobates arenarius Smit & Pešić, 2017
Hygrobates arenarius - Pešić et al. 2017: 1346.
Type locality — The Netherlands, Province of Gelderland, stream at crossing Woestenbergweg and Veenweg (52.336°N, 5.939°E).
BIN — BOLD:AAM2654
BOLD Records — 20 [1 Private]
BIN Distribution — Germany, The Netherlands.
3. Hygrobates corsicus Pešić & Smit, 2017
Hygrobates corsicus - Pešić et al. 2017: 1350.
Type locality — France, Corsica: Ruisseau d'Aitone (42.264°N, 8.846°E).
BIN — BOLD:ADF8471
BOLD Records — 3 [0 Private]
BIN Distribution — France (Corsica).
4. Hygrobates marezaensis Pešić & Dabert, 2017
Hygrobates marezaensis - Pešić et al. 2017: 1354.
Type locality — Montenegro, Podgorica, limnocrenic spring Mareza (42.480°N, 19.182°E).
BIN — BOLD:ADK0646
BOLD Records — 32 [163 Private].
BIN Distribution — Italy, Montenegro, North Macedonia.
5. Hygrobates persicus Pešić & Asadi, 2017
Hygrobates persicus - Pešić et al. 2017: 1358.
Type locality — Iran, Kerman Province, stream in village Sirch (30.198°N, 57.560°E).
BIN — BOLD:ACB5533
BOLD Records — 50 [0 Private].
BIN Distribution — Iran, Turkiye.
New records — Cyprus: Limassol, Mesa Potamos stream (Figure 5A), 34.876°N, 32.910°E, permanent stream, 23 May 2025, leg. Pešić, Tziortzis & Papatheodoulou, 4♀ (sequenced; Table 1); Limnatis (Zygos) stream nr Kouris Dam, intermittent stream, 34.78565°N, 32.94056°E, 23 May 2025, leg. Pešić, Tziortzis & Papatheodoulou, 1♀ (sequenced; Table 1); Paphos district, Kochinas river, 34.877°N, 32.910°E, 20 May 2025 leg. Papatheodoulou, 1♀ (sequenced; Table 1).
Remarks — New for Cyprus.



6. Hygrobates turcicus Pešić, Esen & Dabert, 2017
Hygrobates turcicus - Pešić et al. 2017: 1364.
Type locality — Turkiye, Bayburt Province, Çoruh river (40.206, 40.268).
BINs — BOLD:ADX1364, BOLD:AET3807.
BOLD Records — BOLD:ADX1364: 21 [34 Private]; BOLD:AET3807: 2 [0 Private].
BIN Distribution — BOLD:ADX1364 (Bosnia and Herzegovina, Bulgaria, Turkiye), BOLD:AET3807 (Georgia, Russia).
Remarks — The type specimen of H. turcicus molecularly belongs to BOLD:ADX1364 (Pešić et al. 2017). The genetic distance between the two H. turcicus BINs was estimated at 4.17% (p-distance).
7. Hygrobates grabowskii Pešić, Saboori, Zawal & Dabert, 2019
Hygrobates grabowskii – Pešić et al. 2019b: 2446.
Type locality — Iran, Golestan Province, stream near Shirābād (36.968°N, 55.028°E).
BIN — BOLD:AEG5071, BOLD:AFH9674.
BOLD Records — BOLD:AEG5071: 4 [0 Private]; BOLD:AFH9674: 2 [10 Private].
BIN Distribution — BOLD:AEG5071 (Iran), BOLD:AFH9674 (Turkiye).
Remarks — The type specimen of H. grabowskii molecularly belongs to BOLD:AEG5071 (Pešić et al. 2019b). The genetic distance between the two H. grabowskii BINs was estimated to 6.383 (p-distance).
8. Hygrobates ulii Pešić, Saboori, Zawal & Dabert, 2019
Hygrobates ulii – Pešić et al. 2019b: 2450.
Type locality — Iran, Golestan Province, stream near Shirābād (36.968°N, 55.028°E).
BIN — BOLD:AEG0066.
BOLD Records — 6 [0 Private].
BIN Distribution — Iran.
9. Hygrobates balcanicus Pešić, 2020
Hygrobates balcanicus – Pešić et al. 2020: 1706.
Type locality — Bulgaria, stream below Zemen town (42.469°N, 22.727°E).
BIN — BOLD:AEG3198
BOLD Records — 13 [3 Private].
BIN Distribution — Bulgaria, Portugal, Russia, Serbia.
10. Hygrobates mediterraneus Pešić, 2020
Hygrobates mediterraneus – Pešić et al. 2020: 1710.
Type locality — Montenegro, Bar, Međurječka rijeka stream (42.023°N, 19.220°E).
BIN — BOLD:AED2190
BOLD Records — 3 [6 Private]
BIN Distribution — Greece, Montenegro.
11. Hygrobates cyrnusensis Pešić & Smit, 2023
Hygrobates cyrnusensis – Pešić et al. 2023a: 267.
Type locality — France, Corsica, Rivière La Figarella (42.487°N, 8.805°E).
BIN — BOLD:AEO0837.
BOLD Records — 3 [0 Private].
BIN Distribution — France (Corsica).
12. Hygrobates lusitanicus Pešić & Girão sp. nov.
Type locality — Portugal, Bragança, Torre de Dona Chama, Tuela river (41.665°N, 7.146°W).
BIN — BOLD:AFX1756.
BOLD Records — 6 [0 Private].
BIN Distribution — Portugal.
Discussion
With the discovery of the new species in this study, the Hygrobates fluviatilis complex now includes 12 species. Remarkably, eleven of these have been described since 2017, primarily due to the integration of DNA barcoding, which has refined our understanding of the taxonomic diversity within this complex. All species in the H. fluviatilis complex are represented by more than one barcode in the BOLD database. In total, 13 annotated species names are associated with 787 BOLD records, of which 332 are public and 455 remain private.
With the exception of H. turcicus and H. grabowskii, all other species are associated with a single BIN, which is congruent with our morphological identifications. Both H. turcicus and H. grabowskii are represented by two BINs each. As noted by various authors, the BIN system represents a robust species proxy (Ratnasingham and Hebert 2013) that enables standardized and rapid species identification and diversity assessments.
The BIN clustering and ASAP algorithms yields similar results, with two exceptions where multiple BINs are found, and the results are inconsistent with the final species assignments. Specifically, the ASAP delimitation method fused two BINs (ADX1364 and AET3807) in the case of H. turcicus, while it identified the two BINs of H. grabowskii (AEG5071 and AFH9674) as separate Molecular Operational Taxonomic Units (MOTUs). Recently, Pešić et al. (2026b) emphasized that the relatively high genetic divergence (6.13% p-distance) between the two BINs of H. grabowskii indicates a phylogeographic break (and the possible presence of cryptic species), between these clusters, necessitating further taxonomic investigation.
The occurrence of Hygrobates persicus in Cyprus: biogeographic implications
The finding of Hygrobates persicus in Cyprus highlights an interesting distribution pattern and provides insight into the colonization of the island by mites of the H. fluviatilis complex. A time-calibrated phylogeny recently conducted by Pešić et al. (2023a) revealed that H. persicus diverged during the Late Miocene (8.9–6.3 Ma). This divergence occurred significantly earlier than the final separation of Cyprus from the mainland (south-central Anatolia), which took place during the Pleistocene–Holocene within the last 2.6 million years (Yılmaz 2021). Therefore, it is reasonable to assume that the populations of H. persicus on the island have not had sufficient ''evolutionary time'' to diverge into a distinct species.
The situation is markedly different in Corsica, as demonstrated by Pešić et al. (2023a). This island was colonized twice during different geological periods; in each instance, the lineage became ''trapped'' on the land fragment and evolved in isolation, leading to the presence of two endemic species: H. corsicus and H. cyrnusensis. The estimated time of separation for H. corsicus from the continental ancestors that colonized the Corsico-Sardinian microplate is calculated at approximately 18 Ma (Rosenbaum and Lister 2004). In contrast, the divergence (~7 Ma) between the common ancestor of H. cyrnusensis and its sister mainland lineage, H. fluviatilis, coincides with the splitting of Corsica and Sardinia from Calabria during the opening of the Tyrrhenian Sea, which occurred between 8.6 Ma and 7.6 Ma (see Pešić et al. 2023a for further discussion).
Acknowledgements
This work was partially supported by Biodiversity Genomics Europe (Grant no.101059492) that is funded by Horizon Europe under the Biodiversity, Circular Economy and Environment call (REA.B.3); co-funded by the Swiss State Secretariat for Education, Research and Innovation (SERI) under contract number 22.00173 and 24.00054; and by the UK Research and Innovation Agency under the Department for Business, Energy and Industrial Strategy's Horizon Europe Guarantee Scheme.
SF was funded by the Fundação para a Ciência e a Tecnologia (FCT) through the program'Stimulus of Scientific Employment, Individual Support—3rd Edition'
(https://doi.org/10.54499/2020.03526.CEECIND/CP1601/CP1649/CT0007 ![]() ). The authors would like to acknowledge Lousada Municipality for the logistic support during the fieldwork and Adriana Padilha, Ana Cruz-Oliveira, Luís Guilherme Sousa and Luís P. da Silva for the great company during fieldwork. We thank Reinhard Gerecke (Tübingen) and Joanna Mąkol (Wrocław), whose constructive comments improved this work.
). The authors would like to acknowledge Lousada Municipality for the logistic support during the fieldwork and Adriana Padilha, Ana Cruz-Oliveira, Luís Guilherme Sousa and Luís P. da Silva for the great company during fieldwork. We thank Reinhard Gerecke (Tübingen) and Joanna Mąkol (Wrocław), whose constructive comments improved this work.
Supplementary material
List of specimens of the Hygrobates fluviatilis complex used for building the Neighbor-Joining (NJ) tree (Fig. 1). BINs are based on the barcode analysis from 1 March 2026. Specimens of H. lusitanicus sp. nov. are listed in Table 1.
acarologia_4877_Supplementary_Material.docx
References
- Edgar R.C. 2004. MUSCLE: multiple sequence alignment with high accuracy and high 679 throughput. Nucleic Acids Res., 32(5): 1792-1797. https://doi.org/10.1093/nar/gkh340
- Gerecke R., Gledhill T., Pešić V., Smit H. 2016. Chelicerata: Acari III. In: Gerecke R. (ed.) Süßwasserfauna von Mitteleuropa, Bd. 7/2-3. Springer-Verlag Berlin, Heidelberg, pp. 1-429. https://doi.org/10.1007/978-3-8274-2689-5
- Kimura M. 1980. A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol., 16: 111-120. https://doi.org/10.1007/BF01731581
- Klimov P.B., Stolbov V.A, Kazakov, D.V, Filimonova M.O, Sheykin S.D. 2022. A DNA barcoding and photo-documentation resource of water mites (Acariformes, Hydrachnidia) of Siberia: Accurate species identification for global climate change monitoring programs. Syst. Appl. Acarol., 27(12): 2493-2567. https://doi.org/10.11158/saa.27.12.8
- Kumar S., Stecher G., Li M., Knyaz C., Tamura K. 2018. MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol. Biol. Evol., 35: 1547-1549. https://doi.org/10.1093/molbev/msy096
- Pešić V., Smit H. 2022. Water mites of Corsica: DNA barcode and morphological evidences. Int. J. Acarol., 48: 418-428. https://doi.org/10.1080/01647954.2022.2086619
- Pešić V., Asadi M., Cimpean M., Dabert M., Esen Y., Gerecke R., Martin P., Savić A., Smit H., Stur E. 2017. Six species in one: Evidence of cryptic speciation in the Hygrobates fluviatilis complex (Acariformes, Hydrachnidia, Hygrobatidae). Syst. Appl. Acarol., 22: 1327-1377.
- Pešić V., Broda Ł., Dabert M., Gerecke R., Martin, P., Smit H. 2019a. Re-established after hundred years: Definition of Hygrobates prosiliens Koenike, 1915, based on molecular and morphological evidence, and redescription of H. longipalpis (Hermann, 1804) (Acariformes, Hydrachnidia, Hygrobatidae). Syst. Appl. Acarol., 24(8): 1490-1511. https://doi.org/10.11158/saa.24.8.10
- Pešić V., Saboori A., Zawal A., Dabert M. 2019b. Hidden but not enough: DNA barcodes reveal two new species in Hygrobates fluviatilis complex from Iran (Acariformes, Hydrachnidia, Hygrobatidae). Syst. Appl. Acarol., 24(12): 2439-2459. https://doi.org/10.11158/saa.24.12.11
- Pešić V., Jovanović M., Manović A., Zawal A., Bańkowska A., Ljubomirova L., Karaouzas I., Dabert M. 2020. Molecular evidence for two new species of the Hygrobates fluviatilis complex from the Balkan Peninsula (Acariformes, Hydrachnidia, Hygrobatidae). Syst. Appl. Acarol., 25(9): 1702-1719. https://doi.org/10.11158/saa.25.9.15
- Pešić V., Zawal A., Manović A., Bańkowska A., Jovanović M. 2021a. A DNA barcode library for the water mites of Montenegro. Biodivers. Data J., 9:e78311. https://doi.org/10.3897/BDJ.9.e78311
- Pešić V., Jovanović M., Manović A., Karaouzas I., Smit H. 2021b. New records of water mites from the Balkans revealed by DNA barcoding (Acari, Hydrachnidia). Ecol. Montenegrina, 49: 20-34. https://doi.org/10.37828/em.2021.49.2
- Pešić V., Esen Y., Gerecke R., Goldschmidt T., Mumladze L., Smit H., Zawal A. 2022. Evidence of cryptic speciation in the Hygrobates calliger complex (Acariformes, Hydrachnidia, Hygrobatidae) with the description of two new species. Ecol. Montenegrina, 59: 101-122. https://doi.org/10.37828/em.2022.59.10
- Pešić V., Smit H., Konopleva ES. 2023a. Molecular phylogeny and description of Hygrobates cyrnusensis sp. nov. reveals multiple colonization of Corsica by water mites of the H. fluviatilis complex (Acariformes, Hydrachnidia, Hygrobatidae). Acarologia, 63(1): 262-274. https://doi.org/10.24349/h12f-rs8x
- Pešić V., Esen E., Gülle P., Zawal A., Jovanović M., Rewicz T., Smit H. 2023b. An appraisal of the water mite genus Hygrobates Koch, 1837 from Türkiye, based on morphological and molecular data (Acariformes, Hydrachnidia, Hygrobatidae). Syst. Appl. Acarol., 28(4): 742-754. https://doi.org/10.11158/saa.28.4.10
- Pešić V., Zawal A., Saboori A., Bańkowska A., Cakmak I., Smit H. 2023c. New records of water mites (Acari, Hydrachnidia) from Iran and Türkiye based on morphology and DNA barcodes with description of one new species. Ecol. Montenegrina, 66: 11-29. https://doi.org/10.37828/em.2023.66.2
- Pešić V., Zawal A, Ferreira S., Benitez-Bosco L., Cruz-Oliveira A., Girão D., Padilha A., Turaccio P., Rossini S., Ballini L., Staffoni G., Fratini S., Ciofi C., Iannucci A., Ekrem T., Stur E. 2024a. DNA barcode library of Portuguese water mites, with the descriptions of two new species (Acari, Hydrachnidia). ZooKeys, 1217: 119-171. https://doi.org/10.3897/zookeys.1217.131730
- Pešić V., Smit H., Girão D., Iannucci A., Stur E., Ferreira S. 2024b. New records of aturid water mites from Portugal, with the integrative description of one new species (Acari, Hydrachnidia, Aturidae). Ecol. Montenegrina, 80: 209-219. https://doi.org/10.37828/em.2024.80.19
- Pešić V., Zawal A., Gülle P., Gülle İ., Jovanović M., Bańkowska A., Musielak S., Smit H. 2025a. Water mite diversity from southwestern Türkiye through the lens of the DNA barcodes, with the description of one new species (Acari, Hydrachnidia). ZooKeys, 1232: 205-236. https://doi.org/10.3897/zookeys.1232.142699
- Pešić V., Zawal A., Bańkowska A., Araujo R., Sługocki Ł., Rewicz T., Krakowiak M., Michoński G., Girão D., Silva, L.P. d., Órfão I. Raposeiro P.M., Ballini L., Stryjecki R., Ekrem T., Ferreira, S. 2025b. Exploring the water mite fauna (Acari, Hydrachnidia) of the Madeira archipelago: DNA Barcoding reveals a remarkable species endemicity. Zootaxa, 5621(5), 501-513. https://doi.org/10.11646/zootaxa.5621.5.1
- Pešić V., Bańkowska A., Goldschmidt T., Hårsaker K., Jovanović M., Kaitetzidou E., Krakowiak M., Kozłowska A., Michaloudi, E, Michoński G., Miliša M., Pozojević I., Rewicz T., Rusiniak O., Sobolak K., Szućko I., Stryjecki R., Stur E., Szlauer-Łukaszewska A., Szenejko M., Zawal A. 2025c. Checklist of water mites from the Balkan Peninsula: second supplement, new DNA barcodes and description of a new species. Zootaxa, 5676(1): 1-74.
- Pešić V., Konopleva E.S., Ianucci A., Ferreira S., Stur E. 2025d. Molecular phylogeny reveals three new water mite species of the genus Teutonia Koenike, 1889, from Portugal (Acari, Hydrachnidia, Teutoniidae). Syst. Appl. Acarol., 30(2): 381-401. https://doi.org/10.11158/saa.30.2.14
- Pešić V., Konopleva E.S., Girão D., Vergata C., Sousa, LG., Ferreira, S. 2026a. Molecular phylogeny reveals a new species of the Hygrobates longipalpis complex from Portugal (Acariformes, Hydrachnidia, Hygrobatidae). Ecol. Montenegrina, 94, 84-97. https://doi.org/10.37828/em.2026.94.3
- Pešić V., Bayçelebi E., Zawal A., Bańkowska A., Jovanović M., Kumru S., Baytaşoğlu H., Kalayci G. 2026b. Molecular diversity of water mites of the Turkish eastern Black Sea basin, with the description of two new species (Acari, Hydrachnidia). Zootaxa, 5763(3): 391-406. https://doi.org/10.11646/zootaxa.5763.3.5
- Puillandre N., Brouillet, S., Achaz, G. 2021. ASAP: assemble species by automatic partitioning. Mol. Ecol. Resour., 21(2): 609-620. https://doi.org/10.1111/1755-0998.13281
- Ratnasingham S., Hebert P.D. 2013. A DNA-based registry for all animal species: the barcode index number (BIN) system. PLoS ONE, 8: e66213. https://doi.org/10.1371/journal.pone.0066213
- Rosenbaum G., Lister G.S. 2004. Neogene and Quaternary rollback evolution of the Tyrrhenian Sea, the Apennines, and the Sicilian Maghrebides. Tectonics, 23: 1-17. https://doi.org/10.1029/2003TC001518
- Ström, H. 1768. Beskrivelse over Norske Insecter. Andet Stykke. Det Trondheimske Selskabs Skrifter, 4: 313-370
- Yılmaz Y. 2021. Geological correlation between northern Cyprus and southern Anatolia. Can. J. Earth Sci., 58(6), 640-662. https://doi.org/10.1139/cjes-2020-0129



2026-03-11
Date accepted:
2026-04-09
Date published:
2026-04-14
Edited by:
Mąkol, Joanna

This work is licensed under a Creative Commons Attribution 4.0 International License
2026 Pešić, Vladimir; Girão, Dinis; Tziortzis, Iakovos; Papatheodoulou, Athina and Ferreira, Sónia
Download the citation
RIS with abstract
(Zotero, Endnote, Reference Manager, ProCite, RefWorks, Mendeley)
RIS without abstract
BIB
(Zotero, BibTeX)
TXT
(PubMed, Txt)



