Contribution to the knowledge of Parichoronyssus bakeri Morales-Malacara and Guerrero, 2007 (Mesostigmata: Macronyssidae): new locality and host-association records with additional molecular data
Mello-Oliveira, Victor de Souza  1
; Jacinavicius, Fernando de Castro
1
; Jacinavicius, Fernando de Castro  2
; Guerro-de-Souza, Mariane
2
; Guerro-de-Souza, Mariane  3
; Dornelas Júnior, Leormando Fortunato
3
; Dornelas Júnior, Leormando Fortunato  4
; Castro-Santiago, Ana Carolina
4
; Castro-Santiago, Ana Carolina  5
; Calchi, Ana Cláudia
5
; Calchi, Ana Cláudia  6
; Lourenço, Elizabete Captivo
6
; Lourenço, Elizabete Captivo  7
; Dowling, Ashley P. G.
7
; Dowling, Ashley P. G.  8
; Famadas, Katia Maria
8
; Famadas, Katia Maria  9
; André, Marcos Rogério
9
; André, Marcos Rogério  10
; Camargo, Luis Marcelo Aranha
10
; Camargo, Luis Marcelo Aranha  11
; Barros-Battesti, Darci Moraes
11
; Barros-Battesti, Darci Moraes  12
and Bassini-Silva, Ricardo
12
and Bassini-Silva, Ricardo  13
13
1Vector-Borne Bioagents Laboratory (VBBL), Departamento de Patologia, Reprodução e Saúde Única, Faculdade de Ciências Agrárias e Veterinárias-UNESP, Jaboticabal, SP, Brazil.
2Laboratório de Coleções Zoológicas, Instituto Butantan, São Paulo, SP, Brazil.
3Vector-Borne Bioagents Laboratory (VBBL), Departamento de Patologia, Reprodução e Saúde Única, Faculdade de Ciências Agrárias e Veterinárias-UNESP, Jaboticabal, SP, Brazil.
4Instituto Leônidas e Maria Deane, Fundação Oswaldo Cruz (FIOCRUZ/ILMD), Manaus, Amazonas, Brazil & Instituto de Ciências Biomédicas V, Universidade de São Paulo, São Paulo, Brazil.
5Departamento de Medicina Veterinária Preventiva e Saúde Animal, FMVZ-USP, São Paulo, SP, Brazil.
6Vector-Borne Bioagents Laboratory (VBBL), Departamento de Patologia, Reprodução e Saúde Única, Faculdade de Ciências Agrárias e Veterinárias-UNESP, Jaboticabal, SP, Brazil.
7Laboratório de Ecologia de Mamíferos, Departamento de Ecologia, Universidade do Estado do Rio de Janeiro, Rio de Janeiro, RJ, Brazil.
8Department of Entomology and Plant Pathology, University of Arkansas, Fayetteville, AR, USA.
9Laboratório de Artrópodes Parasitos, Departamento de Parasitologia Animal, Universidade Federal Rural do Rio de Janeiro, Seropédica, RJ, Brazil.
10Vector-Borne Bioagents Laboratory (VBBL), Departamento de Patologia, Reprodução e Saúde Única, Faculdade de Ciências Agrárias e Veterinárias-UNESP, Jaboticabal, SP, Brazil.
11Instituto Leônidas e Maria Deane, Fundação Oswaldo Cruz (FIOCRUZ/ILMD), Manaus, Amazonas, Brazil & Instituto de Ciências Biomédicas V, Universidade de São Paulo, São Paulo, Brazil & Instituto Nacional de Ciência de Tecnologia de Epidemiologia da Amazônia Ocidental (INCT-EPIAMO), Porto Velho, Rondônia, Brazil & Centro de Medicina Tropical de Rondônia (CEPEM) da Secretaria de Estado da Saúde de Rondônia, Monte Negro, Rondônia, Brazil.
12Vector-Borne Bioagents Laboratory (VBBL), Departamento de Patologia, Reprodução e Saúde Única, Faculdade de Ciências Agrárias e Veterinárias-UNESP, Jaboticabal, SP, Brazil.
13✉ Vector-Borne Bioagents Laboratory (VBBL), Departamento de Patologia, Reprodução e Saúde Única, Faculdade de Ciências Agrárias e Veterinárias-UNESP, Jaboticabal, SP, Brazil & Laboratório de Coleções Zoológicas, Instituto Butantan, São Paulo, SP, Brazil.
2024 - Volume: 64 Issue: 1 pages: 56-62
https://doi.org/10.24349/r5x8-stb5Original research
Keywords
Abstract
Introduction
The family Macronyssidae Oudemans, 1936 stands out among mites of medical-veterinary importance since some species can cause dermatitis and even more severe injuries in animals and humans, as well as can carry pathogens, playing a role as putative vectors in the epidemiological cycles of illnesses (Reeves 2007; Hornok et al. 2012; Bassini-Silva et al. 2019; 2022a). Despite that, this family still needs studied since little is known about species diversity, host-parasite associations, biology, and associated microbiome.
Macronyssidae is distributed worldwide and includes 35 genera and over 200 species (Radovsky 2010; Orlova et al. 2017; Bassini-Silva et al. 2021; 2022b). Most species parasitize vertebrates, with approximately 75% of the described species having bats as their primary hosts (Radovsky 2010). In Brazil, 15 genera and 29 species of Macronyssidae have been recorded (Bassini-Silva et al. 2021; 2022b), with about half the diversity recorded parasitizing bats (Bassini-Silva et al. 2021; 2022b).
The genus Parichoronyssus Radovsky, 1966 currently includes 12 species described in New World bats (Morales-Malacara and Guerrero 2020). Four of the 12 species are known to occur on phyllostomid bats in Brazil: Parichoronyssus bakeri Morales-Malacara and Guerrero, 2007; Parichoronyssus crassipes Radovsky, 1967; Parichoronyssus euthysternum Radovsky, 1967; and Parichoronyssus sclerus Radovsky, 1966 (Whitaker and Mumford 1977; Azevedo et al. 2002; Almeida et al. 2011; Moras et al. 2013).
So far, the distribution of these four species is restricted to the southeastern region of Brazil, with P. bakeri, P. crassipes, and P. sclerus only recorded in the state of Minas Gerais (Whitaker and Mumford 1977; Azevedo et al. 2002; Moras et al. 2013) and P. euthysternum in the state of Rio de Janeiro (Almeida et al. 2011). Host records in Brazil are also few, with P. bakeri, P. crassipes, and P. euthysternum collected on the little yellow-shouldered bat, Sturnira lilium (É. Geoffroy, 1810) (Azevedo et al. 2002; Almeida et al. 2011; Moras et al. 2013), P. sclerus on the tailed tailless bat, Anoura caudifer (É. Geoffroy, 1818) (Whitaker and Mumford 1977), and P. bakeri on the Seba's short-tailed bat, Carollia perspicillata (Linnaeus, 1758) (Moras et al. 2013).
This study aimed to identify morphologically and molecularly previously unidentified macronyssid specimens stored in the tissue bank (-80C) at the Acarological Collection of the Butantan Institute (IBSP). Both morphological and molecular data were used to support identification.
Material and methods
The specimens examined in the present study were stored in ethyl alcohol (100%) at (-80C) in the tissue bank at the Acarological Collection of the Instituto Butantan (IBSP). In order to identify these mites, while some specimens were used for DNA extraction, others were slide-mounted in Hoyer's medium, following the protocols by Walter and Krantz (2009). The morphological identification until the genus level followed Radovsky (2010), while the key previously described by Morales-Malacara and Guerrero (2020, 2023) was used for species identification. The original description was consulted to support the identification of the material (Morales-Malacara and Guerrero 2007).
Slide-mounted specimens were examined using a Leica DFC 500 digital camera and a phase-contrast Leica DM4000B optical microscope. Extended focal range images were composed using Leica Application Suite version 2.5.0. The images were prepared with Adobe Photoshop v.13.0 software.
Each mite pulled for DNA analysis was individually placed into a 1.5 µL Eppendorf® microtube. DNA extraction was performed following the manufacturer's QIAGEN DNeasy Blood & Tissue Kit protocols. After DNA extraction, the remaining cuticles were slide-mounted as vouchers and identified following the protocols described above.
Three molecular markers (18S rRNA, 16S rRNA, and COI) were targeted for molecular characterization. The following primer pairs were used to target each gene fragment: 18S rRNA using the primers Mite18S-1F (5′-ATATTGGAGGGCAAGTCTGG-3′) and Mite18S-1R (5′-TGGCATCGTTTATGGTTAG-3′) (Otto and Wilson 2001); 16S rRNA using the primers 16S+1 (5′-CTGCTCAATGATTTTTTAAATTGCTGTGG-3′) and 16S–1 (5′-CCGGTCTGAACTCAGATCAAGT-3′) (Mangold et al. 1998); and COI using the primers LCO1490 (5′-GGTCAACAAATCATAAAGATATTGG-3′) and HCO2198 (5′-TAAACTTCAGGGTGACCAAAAAATCA-3′) (Folmer et al. 1994). PCR reagents concentration and thermal cycler conditions followed the abovementioned works. Negative (ultrapure water type I) and positive controls (pool of Tyrophagus sp.) were used for each reaction. All positive products were purified with ExoSap-IT (GE Healthcare® Pittsburgh, PA).
Sanger sequencing was performed at the ''Centro de Pesquisa sobre Genoma Humano e Células Tronco do Instituto de Biociências″, University of São Paulo, São Paulo State. The obtained sequences were assembled using Sequencing Analysis 5.3.1 software and submitted to BLAST analysis (Altschul et al. 1990) to infer similarities with other mite sequences available in GenBank. Different haplotypes were visually discriminated after an alignment using the CLUSTAL W algorithm (Thompson et al. 1994) implemented in Geneious R11 (Kearse et al. 2012).
Results
A total of 85 macronyssid specimens collected from the greater spear-nosed bat, Phyllostomus hastatus (Pallas, 1767) (Chiroptera: Phyllostomidae) were examined and identified as Parichoronyssus bakeri (Figure 1). Sixty-five of the total number of specimens were used in the molecular analysis, and only three 18S rRNA and six 16S rRNA sequences were obtained from the females of the mites. The obtained sequences were deposited in Genbank as reference sequences for the genus and species. COI was not successfully amplified from any specimen.
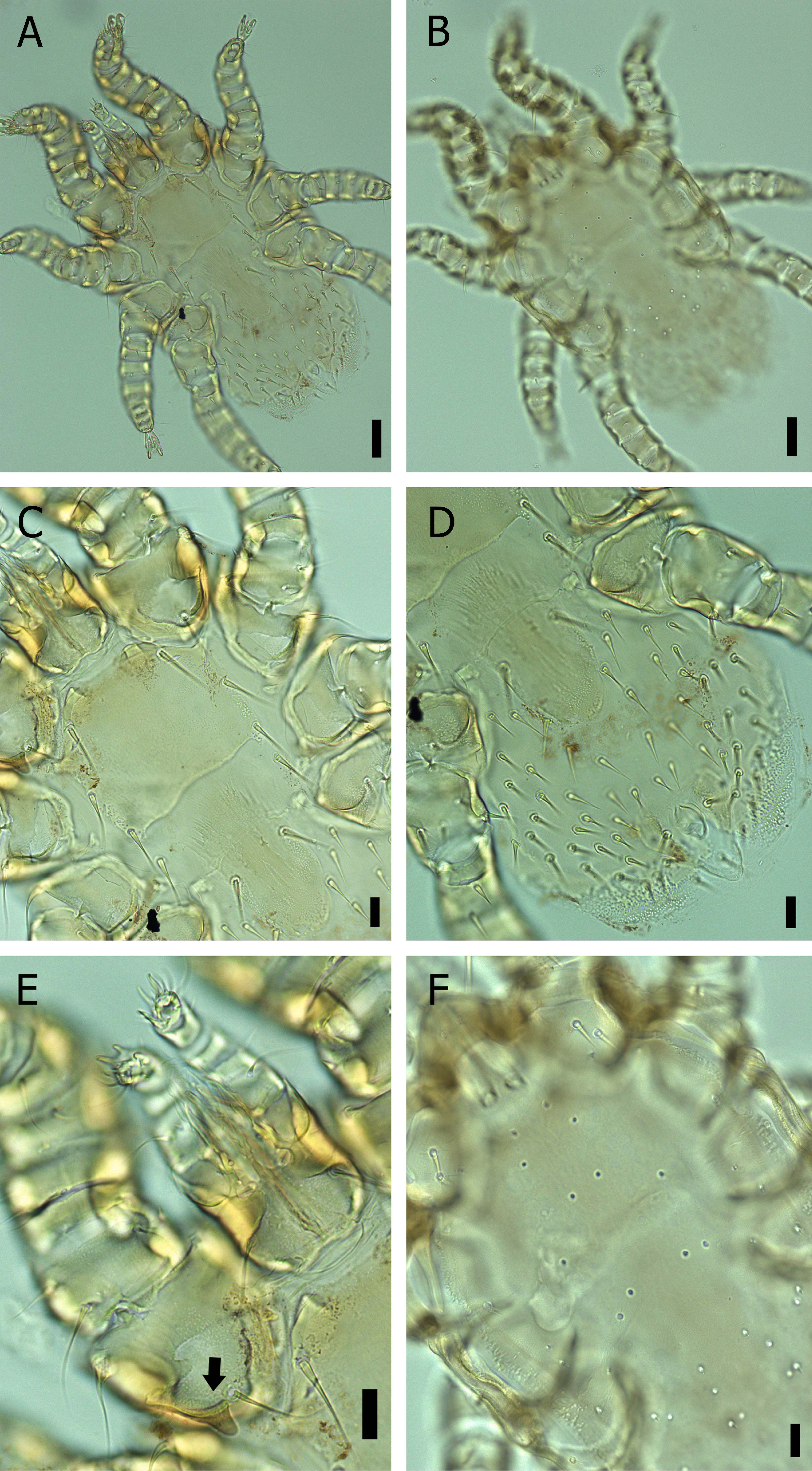


The females of P. bakeri (Figure 1) can be separated from other Parichoronyssus by having the dorsal shield with 13 pairs of setae; trichopores present in the place of setae j4 to j6, and z5; seta r2 absent; more than 24 pairs of setae on the integument of the ventral opisthosoma; entire anal shield without a line interrupting the perianal zone in the anterior region; coxal field I with a spine-like projection with rounded tip in the posterior region (Figure 1E).
Only diagnostic characteristics of the females are being highlighted here since we have yet to obtain sequences of males and nymphal stages (deutonymph and protonymph) to confirm the species. According to the literature, these other stages were morphologically identified, but it is worth noting that different species of Parichoronyssus can parasitize the same host.
The three partial 18S rRNA sequences were obtained from two different lots (IBSP 18475 and 18513) from Rio de Janeiro State and were identical among them, constituting a single haplotype (GenBank access number OR242724). Compared with the sequences available in Genbank, this sequence showed 96.15% identity (query cover: 99%; e-value: 0.0) with Chiroptonyssus haematophagus (GenBank access number: FJ911855). While the six partial 16S rRNA sequences were obtained from three different lots (IBSP 18475, 18513, and 18514), also from the same state. Five sequences were identical between them, constituting a single haplotype, and the other sequence with a difference when compared to the haplotype (GenBank access number OR242725 and OR242726). The two haplotypes of 16S rRNA partial sequences showed 89.44% and 88.89% identity (query cover: 98% and 99%; e-value: 6e-14 and 4e-137, respectively) with Chiasmanyssus cavernicola (GenBank access number: MW522530).
Material examined. BRAZIL — 14♀ and 8♂ (IBSP 18370); Monte Negro Municipality, Rondônia State (63°14′2″W, 10°17′42″S); 20 May 2021; ex. Phyllostomus hastatus (Pallas, 1767) (Chiroptera: Phyllostomidae); Dornelas L. F. Jr. coll. 8♀ and 4♂ (IBSP 18376); same data. 14♀, 2♂ and 7PN (IBSP 18475); Mário Takume Farm, Tinguá Neighborhood, Nova Iguaçu Municipality, Rio de Janeiro State; 17 Nov. 2011; same host; Lourenço E.C. et al. coll. 5♀ and 12PN (IBSP 18514); same locality and host; 14 Apr. 2012. 11♀ (IBSP 18513); same locality and host; 17 Nov. 2011.
Discussion
Four species of Parichoronyssus have been recorded from Brazil: P. bakeri, P. crassipes, P. euthysternum, and P. sclerus. Based on known host records in the Neotropics (Azevedo et al. 2002; Morales-Malacara and Guerrero 2020), the previous reports from Brazil for some of these species might need to be revised. This might have happened due to possible contamination during collection and handling, accidental transmission of mites to the wrong hosts, or even misidentification. For example, the records of P. bakeri from C. perspicillata and S. lilium in Minas Gerais State (Moras et al. 2013) are problematic because elsewhere, this species has only been associated with the genus Phyllostomus. Additionally, P. crassipes from Sturnira lilium (Azevedo et al. 2002) is also questionable since P. crassipes is a mite species that primarily parasitizes Carollia spp. Specimens from those studies should be reexamined to validate species identification and document the natural distribution of these species in the Brazilian territory.
Regarding the molecular analysis, the only Parichoronyssus sequence available in Genbank is for a fragment of the 28S rRNA (Dowling and O'Connor 2010) without species identification. The present study contributed to future molecular identification of Parichoronyssus spp. by providing sequences for two molecular markers, namely 18S rRNA and 16S rRNA, aiming at improving the genetic representation of P. bakeri in Genbank. Also, we provide diagnostic images and additional information about P. bakeri females, as well as new locality records for this species in Rio de Janeiro and Rondônia States, all associated with Ph. hastatus.
Acknowledgments
To Gabrielle Ribeiro de Andrade, Maria Cristina Ferreira do Rosário from the Laboratório de Coleções Zoológicas, Instituto Butantan, for technical contribution; To Valeria Castilho Onofrio for giving access to Acarological Collection of the Instituto Butantan. To Juan Bibiano Morales-Malacara for kindly sending the micrographs of the type series of P. bakeri. This work was supported by the Conselho Nacional de Desenvolvimento Científico e Tecnológico under the Grant CNPq no. 402575/2021-0 (FCJ), the Productivity Grant to MRA (CNPq Process #303701/2021-8), and DMB-B (CNPq Process #303802/2021-9), as well as the Fundação de Amparo à Pesquisa do Estado de São Paulo under Grant FAPESP no. 2023/06748-9 (MG-S), 2023/06878-0 (VSM-O), 2022/08620-7 (VSM-O), 2019/19853-0 (FCJ), 2021/06758-9 (ACC-S), 2020/07826-5 (ACC), 2017/01416-7 (RB-S), 2018/24667-8 (RB-S), and 2020/11755-6 (RB-S); and also, was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES) - Finance Code 001.
References
- Almeida J.C., Silva S.S.P., Serra-Freire N.M., Valim M.P. 2011. Ectoparasites (Insecta and Acari) associated with bats in Southeastern Brazil. J. Med. Entomol., 48: 753-757. https://doi.org/10.1603/ME09133
- Altschul S.F., Gish W., Miller W., Myers E.W., Lipman D.J. 1990. Basic local alignment search tool. J. Mol. Biol., 215: 403-410. https://doi.org/10.1016/S0022-2836(05)80360-2
- Azevedo A.A., Linardi P.M., Coutinho M.T.Z. 2002. Acari Ectoparasites of Bats from Minas Gerais, Brazil. J. Med. Entomol., 39(3): 553-555. https://doi.org/10.1603/0022-2585-39.3.553
- Bassini-Silva R., Jacinavicius F.C., Hernandes F.A., Ochoa R., Bauchan G.R., Dowling A.P.G., Barros-Battesti D.M. 2019. Dermatitis in humans caused by Ornithonyssus bursa (Berlese 1888) (Mesostigmata: Macronyssidae) and new records from Brazil. Rev. Bras. Parasitol. Vet., 28: 134-139. https://doi.org/10.1590/s1984-296120180097
- Bassini-Silva R., Jacinavicius F.C., Huang-Bastos M., Dowling A.P.G, Barros-Battesti D.M. 2021. A checklist of macronyssid species (Mesostigmata: Macronyssidae) from Brazil. J. Med. Entomol., 58: 625-633. https://doi.org/10.1093/jme/tjaa245
- Bassini-Silva, R., Castro-Santiago, A.C., Calchi, A.C., Perles L., Takatsu J.C., Alencar I.D.C.C., Ochoa R., Dowling A.P.G., Werther K., André M.R., Barros-Battesti D.M., Jacinavicius, F.C. 2022a. Sleeping with the enemy: case reports of Ornithonyssus bursa (Berlese, 1888) (Mesostigmata: Macronyssidae) causing human dermatitis in Brazil. Parasitol. Res., 121: 2641-2649. https://doi.org/10.1007/s00436-022-07589-2
- Bassini-Silva R., Jacinavicius F.C., Castro-Santiago A.C., Dowling A.P.G., André M.R., Linardi P.M., Barros-Battesti D.M. 2022b. Occurrence of the mite Acanthonyssus proechimys (Mesostigmata: Macronyssidae) on a spiny rat in Brazil. Acta Amazon., 52: 328-330. https://doi.org/10.1590/1809-4392202200512
- Dowling A.P.G., O'Connor B.M. 2010. Phylogeny of Dermanyssoidea (Acari: Parasitiformes) suggests multiple origins of parasitism. Acarologia, 50: 113-129. https://doi.org/10.1051/acarologia/20101957
- Folmer O., Black M., Hoeh W., Lutz R., Vrijenhoek R. 1994. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol., 3: 294-299.
- Hornok S., Kovács R., Meli M.L., Gönczi E., Hofmann-Lehmann R., Kontschán J., Gyuranecz M., Dá A., Molnár V. 2012. First detection of bartonellae in a broad range of bat ectoparasites. Vet. Microbiol., 159(3-4): 541-543. https://doi.org/10.1016/j.vetmic.2012.04.003
- Kearse M., Moir R., Wilson A., Stones-Havas S., Cheung M., Sturrock S., Buxton S., Cooper A., Markowitz S., Duran C., Thierer T., Ashton B., Meintjes P., Drummond A. 2012. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinform., 28: 1647-1649. https://doi.org/10.1093/bioinformatics/bts199
- Mangold A.J., Bargues M.D., Mas-Coma S. 1998. Mitochondrial 16S rDNA sequences and phylogenetic relationships of species of Rhipicephalus and other tick genera among Metastriata (Acari: Ixodidae). Parasitol. Res., 84(6): 478-484. https://doi.org/10.1007/s004360050433
- Morales-Malacara J.B., Guerrero R. 2007. A new species of Parichoronyssus (Acari: Dermanyssoidea: Macronyssidae) from bats of the genus Phyllostomus (Chiroptera: Phyllostomidae) in Peru and Venezuela, with keys to the species of Parichoronyssus. J. Med. Entomol., 44(1), 8-13. https://doi.org/10.1603/0022-2585(2007)44%5B8:ANSOPA%5D2.0.CO;2
- Morales-Malacara J.B., Guerrero R. 2020. Two new species and new records of mites of the genus Parichoronyssus (Acari: Macronyssidae) from South American bats (Chiroptera), with a key to the known species of the genus. J. Med. Entomol., 57(2): 404-417. https://doi.org/10.1093/jme/tjz196
- Morales-Malacara J.B., Guerrero R. 2023. Un nuevo Parichoronyssus Radovsky, 1966 (Acari: Macronyssidae) asociado con el Murciélago, Rhinophylla fischerae Carter, 1966 (Chiroptera: Phyllostomidae) de Perú, con clave de identificación. Dugesiana, 30(2): 203-208. https://doi.org/10.32870/dugesiana.v30i2.7294
- Moras L.M., Bernardi L.F.O., Graciolli G., Gregorin R. 2013. Bat flies (Diptera: Streblidae, Nycteribiidae) and mites (Acari) associated with bats (Mammalia: Chiroptera) in a high-altitude region in southern Minas Gerais, Brazil. Acta Parasitol., 58: 556-563. https://doi.org/10.2478/s11686-013-0179-x
- Orlova M.V., Kazakov D.V., Orlov O.L., Mishchenko V.A., Zhigalin A.V. 2017. The first data on the infestation of the parti-coloured bat, Vespertilio murinus (Chiroptera, Vespertilionidae), with gamasid mites, Steatonyssus spinosus (Mesostigmata, Gamasina, Macronyssidae). Russ. J. Theriol., 16: 66-73. https://doi.org/10.15298/rusjtheriol.16.1.06
- Otto J.C., Wilson K.J. 2001. Assessment of the usefulness of ribosomal 18S and mitochondrial COI sequences in Prostigmata phylogeny. In: Proctor H.C., Norton R.A., Colloff M.J. editors. Acarology: Proceedings of the 10th International Congress. Melbourne: CSIRO Publishing. p. 100-109.
- Radovsky F.J. 2010. Revision of genera of the parasitic mite family Macronyssidae (Mesostigmata: Dermanyssoidea) of the World, Vol. 170. West Bloomfield, MI, USA: Indira Publishing House, 160 pp.
- Reeves W.K., Loftis A.D., Szumlas D.E., Abbassy M.M., Helmy I.M., Hanafi H.A., Dasch G.A. 2007. Rickettsial pathogens in the tropical rat mite Ornithonyssus bacoti (Acari: Macronyssidae) from Egyptian rats (Rattus spp.). Exp. Appl. Acarol., 41(1-2): 101-107. https://doi.org/10.1007/s10493-006-9040-3
- Thompson J.D., Higgins D.G., Gibson T.J. 1994. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positions-specific gap penalties and weight matrix choice. Nucleic Acids Research, 22: 4673-4680. https://doi.org/10.1093/nar/22.22.4673
- Walter D.E., Krantz G.W. 2009. Chapter 7 - collecting, rearing, and preparing specimens. In: Krantz G.W., Walter D.E. (eds). A manual of acarology. Lubbock: Tech University Press; p. 83-96.
- Whitaker Jr. J.O., Mumford R.E. 1977. Records of ectoparasites from Brazilian mammals. Entomol. News 88: 255-258.



2023-07-08
Date accepted:
2023-12-04
Date published:
2023-12-14
Edited by:
Faraji, Farid

This work is licensed under a Creative Commons Attribution 4.0 International License
2024 Mello-Oliveira, Victor de Souza; Jacinavicius, Fernando de Castro; Guerro-de-Souza, Mariane ; Dornelas Júnior, Leormando Fortunato; Castro-Santiago, Ana Carolina; Calchi, Ana Cláudia; Lourenço, Elizabete Captivo; Dowling, Ashley P. G.; Famadas, Katia Maria; André, Marcos Rogério; Camargo, Luis Marcelo Aranha; Barros-Battesti, Darci Moraes and Bassini-Silva, Ricardo
Download the citation
RIS with abstract
(Zotero, Endnote, Reference Manager, ProCite, RefWorks, Mendeley)
RIS without abstract
BIB
(Zotero, BibTeX)
TXT
(PubMed, Txt)



